---
title: "AxB ANOVA"
author: "Lesson 5"
output:
html_document:
css: http://www.bradthiessen.com/batlab2.css
highlight: pygments
theme: spacelab
toc: yes
toc_depth: 5
fig_width: 5.6
fig_height: 4
---
```{r message=FALSE, echo=FALSE}
# Load the mosaic package
library(mosaic)
```
*****
## Scenario: Cholesterol, Drug, and Obesity
Does a drug designed to lower cholesterol levels actually work?

To study this, researchers randomly select 40 adult males with high cholesterol and randomly assign them to one of two groups:
• 20 subjects receive the **drug**
• 20 subjects receive a **placebo**
After a 6-month regimen of taking the drug or placebo, the subjects' cholesterol levels were measured.
*****
## Data and analysis
Here's a summary of the cholesterol levels of the 40 subjects:
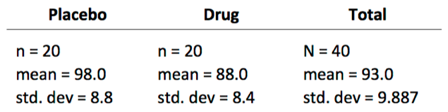
With this data, you could calculate a test statistic for an independent samples t-test (assuming equal variances:
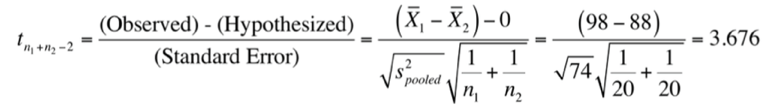
with
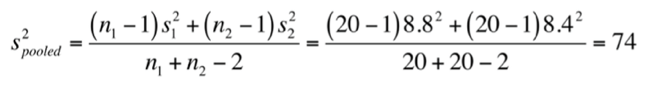
and a critical value of $t_{df=38}^{\alpha =0.025}=2.024$.
1. We could have also chosen to conduct an ANOVA to compare the group means. Based on the results of the t-test, fill-in-the-blanks to complete the ANOVA summary table. What conclusions can we make from the ANOVA or t-test?
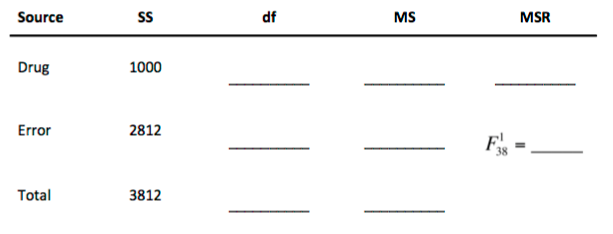
You publish the results of this study and receive the following feedback:
> It is widely known that obese individuals have higher cholesterol than those who are non-obese. Because your *drug* group could have simply had more obese subjects than your *placebo* group, I question the results of your study.
2. Uh oh - you forgot to measure the body mass of your subjects in this study. Is this a legitimate concern?
*****
## Potential datasets (including obesity)
We're going to learn how to conduct an ANOVA when we have 2 independent variables (drug and obesity) and a single dependent variable (cholesterol). When we're able to randomly assign subjects to levels of each independent variable, we have an **AxB ANOVA** (a.k.a. a *two-way ANOVA* or *factorial design*).
Before we get into the concepts and calculations, let's look at some hypothetical results from our cholesterol study.
3. For each scenario displayed below, sketch a plot of the mean cholesterol levels for each combination of obesity and drug. These are sometimes called *interaction plots*
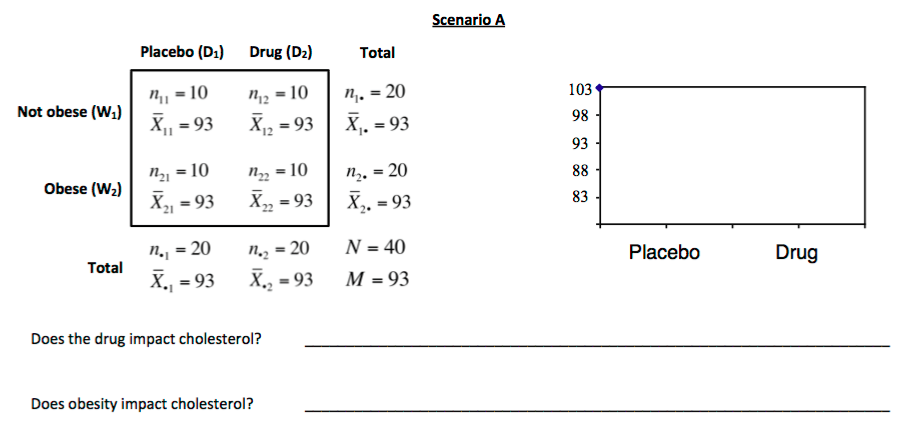
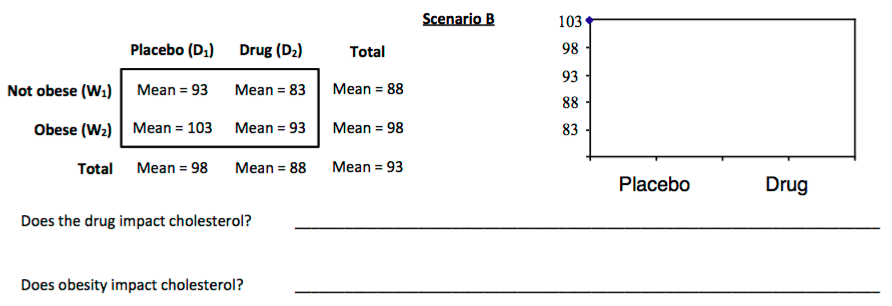
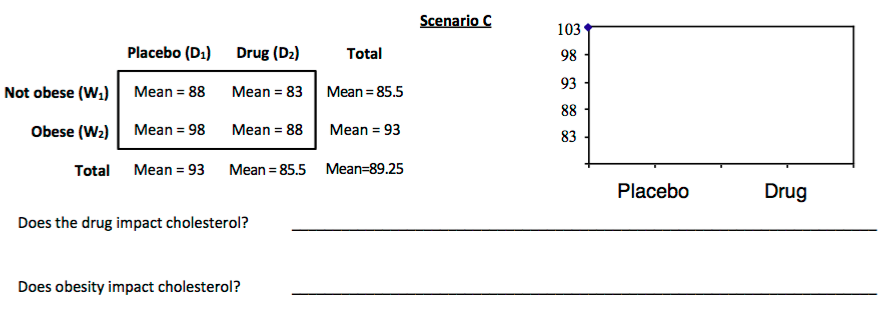
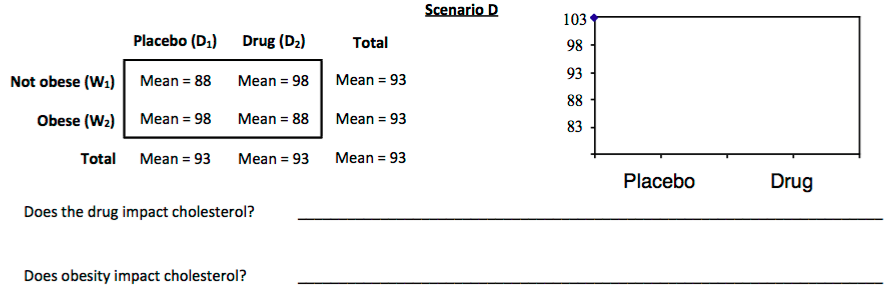
In an AxB ANOVA, we'll estimate two classes of effects: **main effects** and **interaction effects**.
A main effect for a factor (one independent variable) is the overall effect it has on the dependent variable consistently across the levels of the other factor (other independent variable).
In this scenario, for example: **The main effect of the drug is the effect it has on cholesterol consistently for obese and non-obese individuals.**
4. Go back to scenarios A-D and calculate the main effect for the drug and the main effect for obesity. Then, calculate the sum of those effects.
Scenario |Drug effect |Obesity effect |Sum of effects |
---------:|-----------:|--------------:|--------------:|
A :| | | |
B :| | | |
C :| | | |
D :| | | |
The other type of effect we'll estimate is the **interaction effect**. Interaction is present when the two independent variables in combination produce effects that are different from the sum of their main effects.
**Interaction means the effect of one independent variable depends on the other independent variable.** In this study, an interaction effect would mean the effect of the drug on cholesterol depends on the obesity of the subject.
In **Scenario B**, we calculated the main effects for the drug and obesity to each be 5. The sum of those effects is 10. Suppose we take a subject with an average cholesterol level of 93 (the grand mean). If we have no interaction, then we'd expect that subject to have a cholesterol level of 83 (10 points lower) if that subject was given the drug and was non-obese. This is what happens in this scenario with **no interaction** (and we can see this in the **parallel** lines in our means plot).
In **Scenario D**, the main effects were both zero. We'd expect a non-obese subject given the drug would have the same cholesterol level as the grand mean. That's not what the results in the scenario show. The combination of the effects of our independent variables produces an effect different from what we expect from each independent variable separately. We can tell we have interaction because the lines in our means plot are **not parallel**.
5. Is there an interaction effect in Scenario C? Explain.
*****
## Model: Main effects and interaction
6. Imagine you are a subject in this study. What factors could influence your cholesterol level at the end of the 6-month study?
We can write the formal model as: $y_{\textrm{ijk}}=\mu +\alpha _{j}+\beta _{k}+\alpha \beta _{jk}+\epsilon _{ijk}$,
where: $\alpha _{j}=\mu_{Aj}-\mu, \ \ \ \beta _{k}=\mu_{Bk}-\mu, \ \ \ \alpha \beta _{jk}=\mu_{AjBk}-\mu_{Aj}-\mu_{Bk}+\mu, \ \ \ \epsilon _{ijk}=y_{ijk}-\mu_{AjBk}$
7. Show this model is a tautology.
The following table displays the data from this cholesterol study.
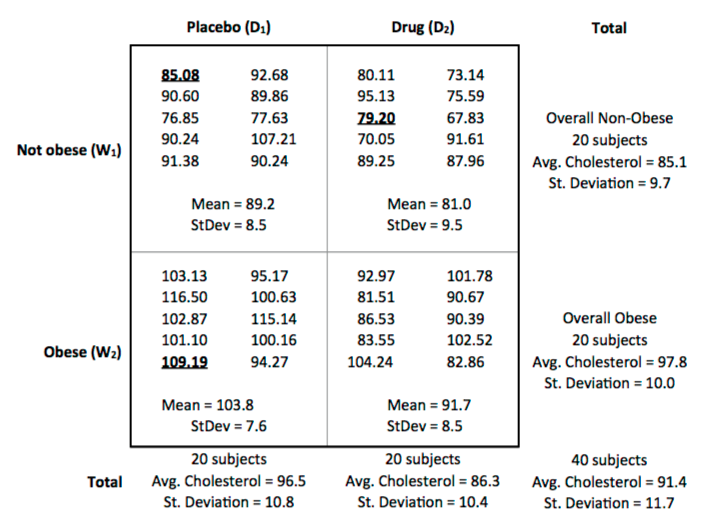
8. The first subject in the table has been highlighted. According to our model, why did this subject end the study with a cholesterol level of 85.08? What are the values of $\alpha _{j}$, $\beta _{k}$, $\alpha \beta _{jk}$, and $\epsilon _{ijk}$?
Let's partition the total variation in cholesterol levels of the subjects in this study.
9. Explain each step in the following derivation:
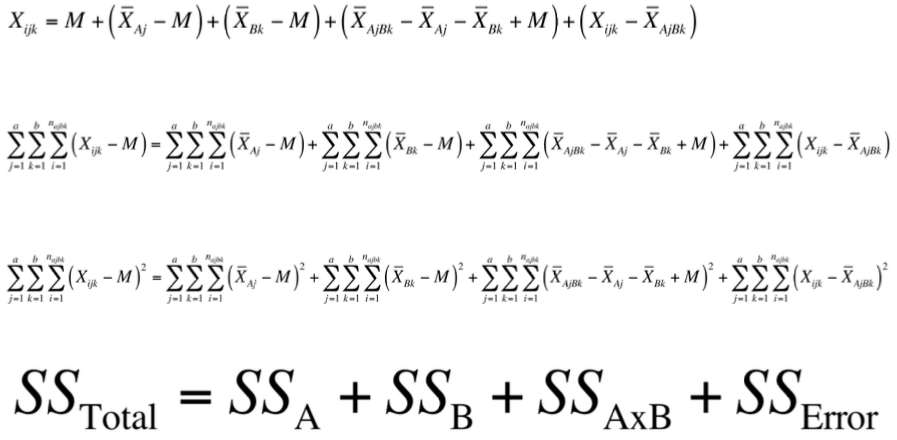
10. Explain what each of those SS values represents.
Once we calculate the SS values, we'll convert them to MS values by dividing by their degrees of freedom. Then, we'll compare the mean squares by taking ratios.
11. Out of all our MS values (MSdrug, MSobesity, MSinteraction, MSerror), which ratio should we calculate to determine if the drug had a non-zero effect on cholesterol? Which ratio would measure the effect of obesity on cholesterol? Which ratio would test for interaction? Which ratio should we calculate first?
Here's a visual display of how the total variation is partitioned in a (one-way) ANOVA and an AxB ANOVA:
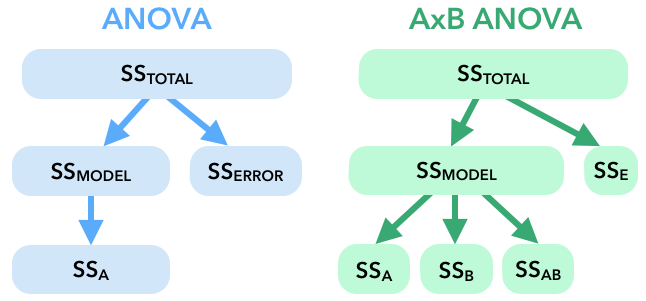
12. Before we calculate our SS, df, and MS values, let's plot the means from the actual data in this study to predict whether we'll find a significant interaction effect.
### Calculating AxB ANOVA summary table
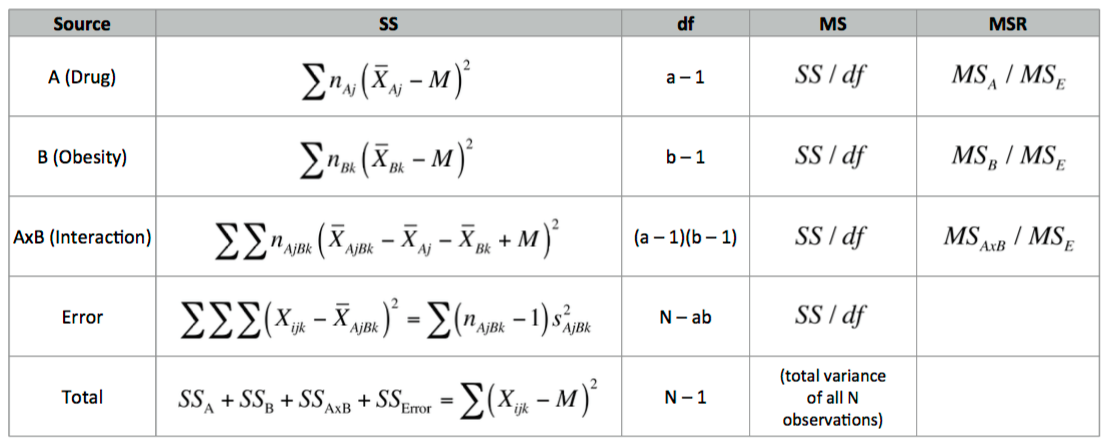

I used R to get the following calculations:
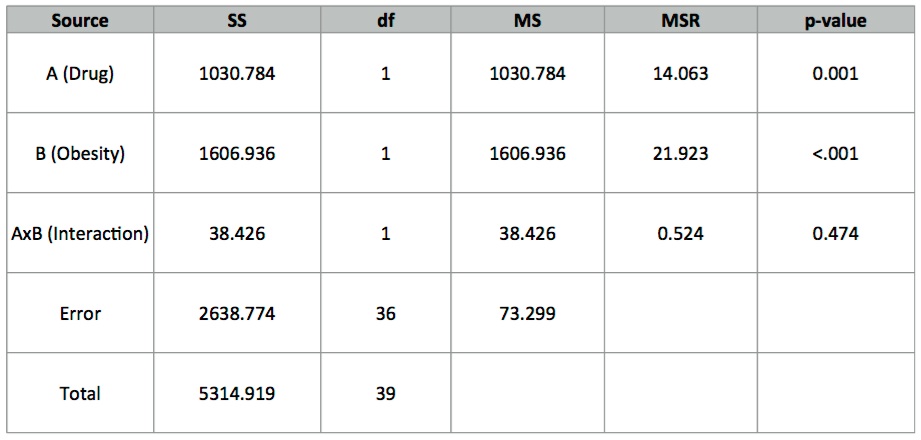
13. Verify the SS and df values. How did R estimate those p-values? Calculate an effect size and state your conclusion(s).
Here's how I calculated this in R:
```{r}
# Load data as "chol1"
chol1 <- read.csv("http://www.bradthiessen.com/html5/data/cholesterol1.csv")
# Construct the model: y ~ x1 + x2 + x1x2
# In R, we use: y ~ x1 * x2 (or: y ~ x1 + x2 + x1:x2)
model1 <- aov(cholesterol ~ drug * obese, data=chol1)
# Summarize results (results differ because of rounding errors)
anova(model1)
```
After conducting the AxB ANOVA, we should check our assumptions/conditions:
```{r, message=FALSE}
# Construct the model again, this time with "lm" instead of "aov"
# There's really no difference except how R stores the results
model1 <- lm(cholesterol ~ drug * obese, data=chol1)
# Construct some plots of the model
mplot(model1)
# Construct one more plot of the model
plotModel(model1)
```
14. Interpret each plot. We'll see these in the next unit (and discuss them in much greater detail).
*****
### Interaction Effects
Consider the same study with slightly different results. I changed the data for the **non-obese, drug** group:
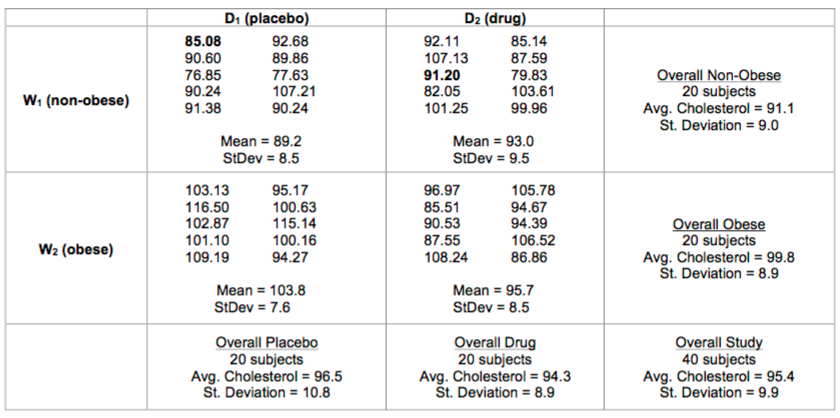
Let's load this data into R:
```{r}
# Load the data as chol2
chol2 <- read.csv("http://www.bradthiessen.com/html5/data/cholesterol.csv")
# Convert the treatment variables into factors
chol2$obese <- as.factor(chol2$obese)
chol2$drug <- as.factor(chol2$drug)
chol2$Group <- as.factor(chol2$Group)
# Let's look at the structure of our dataset
str(chol2)
# Let's look at the distribution of cholesterol in each group
densityplot(~cholest | drug * obese, data=chol2,
main="Cholesterol by drug and obesity",
xlab="Cholesterol",
layout=c(2,2))
```
15. Based on the following interaction plot, do you think we'll find a significant interaction effect?
```{r}
# Here's the syntax for an interaction plot
# x.factor = the variable to be used as the x-axis
# trace.factor = the variable to represent the lines
# response = the dependent variable
interaction.plot(x.factor = chol2$drug, trace.factor = chol2$obese,
response = chol2$cholest, trace.label = c("Obesity"),
xlab = "Treatment (0 = placebo, 1 = drug)",
ylab = "Mean cholesterol level",
col = "steelblue", lw=4)
```
16. Based on the following output, evaluate the conditions necessary to conduct an AxB ANOVA.
```{r}
# Levene's test for homogeneity of variances
leveneTest(cholest ~ drug * obese, data=chol2)
# Shapiro-Wilk's test for normality (one for each of our 4 groups)
grp1 <- round(shapiro.test(chol2$cholest[chol2$Group=="1"])$p.value,3)
grp2 <- round(shapiro.test(chol2$cholest[chol2$Group=="2"])$p.value,3)
grp3 <- round(shapiro.test(chol2$cholest[chol2$Group=="3"])$p.value,3)
grp4 <- round(shapiro.test(chol2$cholest[chol2$Group=="4"])$p.value,3)
# Paste the p-values from the Shapiro-Wilk tests
paste("Group 1: p =", grp1, ". Group 2: p =", grp2, ". Group 3: p =", grp3, ". Group 4: p =", grp4)
```
Let's run the AxB ANOVA and take a look at the summary table:
```{r}
# Run the ANOVA model: y ~ a * b
model2 <- aov(cholest ~ drug * obese, data=chol2)
# Print the summary table
anova(model2)
```
17. What conclusions can we make? Do we have a significant interaction effect? Sketch and label a diagram to display the partitioning of the total variation in cholesterol.
18. State what the interaction effect represents. Based on that explanation, what conclusions can we make regarding the effectiveness of the drug or obesity?
Let's see some of the model plots:
```{r}
# Print model plots
model2 <- lm(cholest ~ drug * obese, data=chol2)
mplot(model2)
```
#### Simple Effects
A significant interaction means the effect of the drug on cholesterol depends on obesity. Because of this, it doesn't make sense to think about the "drug effect" without considering obesity at the same time.
When we find a significant interaction, we can split our data and investigate the **simple effects**.
Because we're most interested in the effect of the drug, let's split our data into obese and non-obese groups. To do this in R, we can use:
```{r}
# Select all data where obese == 0 in the chol2 data.frame
# Store the results in a data.frame called "nonobese"
# The blank space to the right of the comma in "0, ]"
# tells R to select ALL the rows of data
nonobese <- chol2[chol2$obese==0, ]
# Select all data where obese == 1
obese <- chol2[chol2$obese==1, ]
# Note that we could separate our data using dplyr
# The syntax is a little easier to read from left-to-right
# nonobese <- chol2 %>% filter(obese==0)
# obese <- chol2 %>% filter(obese==1)
```
Here's a summary of those split datasets:
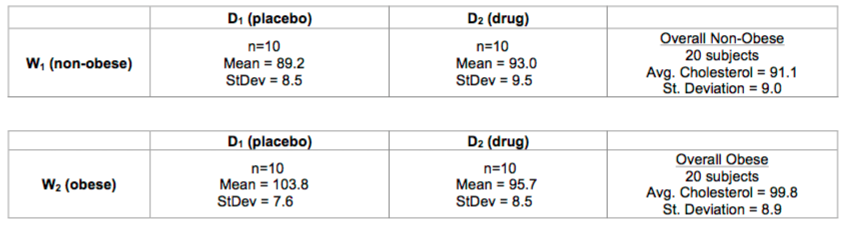
19. Look at the table for the non-obese groups. If we want to compare the mean of the drug group to the mean of the placebo group, what could we do?
Let's conduct t-tests to compare the drug and placebo group means. First, we'll run the test for the non-obese subjects; then we'll run the test for obese subjects:
```{r}
# I'll run a t-test without the equal variance assumption
# Here's the test for nonobese subjects
t.test(cholest ~ drug, data=nonobese, var.equal=FALSE, alternative = c("less"))
# t-test for obese subjects
# Note that we could run an ANOVA to compare these 2 groups, too.
t.test(cholest ~ drug, data=obese, var.equal=FALSE, alternative = c("greater"))
```
20. Based on these simple effects, what conclusions can we make?
*****
## Resampling-based methods
### Randomization-based AxB ANOVA
In conducting the AxB ANOVA (and the t-tests of the simple effects), we had to assume normality, independence, and equal variances. If these assumptions seem unreasonable, we can use randomization-based methods to test for interaction and main effects.
Remember the logic behind randomization-based methods:
- First, we calculate a test statistic of interest.
- We then assume a randomization hypothesis is true.
- Assume the drug, obesity, and their interaction have no effect on cholesterol.
- We can then pretend to go back in time and randomly assign subjects to drug and obesity groups.
- Randomly assigning subjects to obesity groups is impossible, so we should generalize carefully.
- For each randomization of our data, we calculate our test statistic.
- Once we've run 10,000 randomizations, we can determine the likelihood of observing our actual test statistic if the randomization hypothesis were true.
We're going to need to conduct randomization-based tests separately for the interaction and main effects. We'll test for interaction first.
#### Randomization test for interaction
Let's look one more time at our AxB ANOVA output:
```{r}
anova(model2)
```
21. From that output, choose a test statistic that measures the size of the interaction effect?
We'll now randomize the interaction effect by randomly assigning subjects to drug and obesity groups:
```{r}
# I'm going to load the "broom" package. It allows us to easily store
# and access results from a model.
library(broom)
# First, we'll re-run the model
aovmodel <- aov(cholest ~ drug * obese, data=chol2)
# Now we'll use the "tidy" command to store those results
tidy(aovmodel)
# We want to store the 3rd statistic (F-value for interaction = 4.8465)
# We'll store this value as "obs_F_interaction"
obs_F_interaction<- tidy(aovmodel)$statistic[3]
# Now let's run 10,000 randomizations, each time storing the F-value
# We will shuffle the interaction term
random_interact <- do(10000) *
tidy(aov(cholest ~ drug + obese + shuffle(drug):shuffle(obese),
data=chol2))$statistic[3]
# We'll plot a histogram of the F values from these 10,000 randomizations
histogram(~random_interact, data = random_interact,
xlim=c(-.1, 6), xlab = "F-statistic for interaction", width=.05,
col="steelblue", border="white")
# Now, let's get the p-value by counting the proportion of randomization
# F-values greater than our observed test statistic
tally(~ (random_interact >= obs_F_interaction), format="prop", data=random_interact)
# We could have done this without the broom package
# All the extra brackets to the right are there only to
# extract the F statistic for the interaction effect.
# Notice that the F for interaction is the 3rd F in the summary table.
# Therefore, to extract it, I need [[1]][["F value"]][[3]]
# We'll first get our test statistic (F for interaction)
## observed_model <- do(1) *
## summary(aov(cholest ~ drug*obese, data=chol2))[[1]][["F value"]][[3]]
## obs_F_interaction <- observed_model$"[["
# Now let's take 10,000 randomizations
# We'll randomly shuffle the interaction term and get the F value
## random_interact <- do(10000) * summary(aov(cholest ~ drug + obese + shuffle(drug):shuffle(obese),
## data=chol2))[[1]][["F value"]][[3]]
```
22. From this, what conclusion can we draw?
Now that we've found a significant interaction effect, we could split our data and conduct randomization-based tests separately for the obese and non-obese groups.
Let's, instead, pretend as though our interaction effect was **not** significant. If that were the case, we could examine the main effects of the drug and obesity. We can do this via randomization-based tests:
```{r}
# We want to store the 1st statistic (F-value for interaction = 0.632)
# We'll store this value as "obs_F_drug"
obs_F_drug<- tidy(aovmodel)$statistic[1]
# Now let's run 10,000 randomizations, each time storing the F-value
# We will shuffle the interaction term
random_drug <- do(10000) *
tidy(aov(cholest ~ shuffle(drug) + obese + drug:obese,
data=chol2))$statistic[1]
# We'll plot a histogram of the F values from these 10,000 randomizations
histogram(~random_drug, data = random_interact,
xlim=c(-.1, 10), xlab = "F-statistic for interaction", width=.05,
col="steelblue", border="white")
# Now, let's get the p-value by counting the proportion of randomization
# F-values greater than our observed test statistic
tally(~ (random_drug >= obs_F_drug), format="prop", data=random_interact)
```
23. Does this p-value agree with the p-value from the AxB ANOVA?
I won't run the following code, but it would test the obesity main effect:
```{r, eval=FALSE}
# Obesity effect
observed_model <- do(1) *
summary(aov(cholest ~ drug*obese, data=chol2))[[1]][["F value"]][[2]]
obs_F_obesity <- observed_model$"[["
# Shuffle assignment to obesity groups
random_obesity <- do(1000) * summary(aov(cholest ~ drug + shuffle(obese) + drug:obese, data=chol2))[[1]][["F value"]][[2]]
# Histogram
histogram(~random_obesity, data = random_obesity, xlim=c(-.1, 15),
xlab = "F-statistic for obesity effect", width=.1, col="steelblue", border="white")
# p-value
tally(~ (random_obesity >= obs_F_obesity), format="prop", data=random_interact)
```
### Bootstrap CI for effect size
We've calculated an effect size of `eta-squared = 0.304`. Suppose we wanted to construct confidence interval around that eta-squared. We can do that with bootstrap methods:
```{r}
# Take 10,000 bootstrap samples (with replacement) and calculate the AxB ANOVA
bootmodel <- do(10000) * lm( cholest ~ drug * obese, data=resample(chol2))
# Store the eta-squared values as "bootetasquared"
bootetasquared <- bootmodel$r.squared
# Plot the eta-squared values from our bootstrap samples
densityplot(~bootetasquared, plot.points = FALSE, col="steelblue", lwd=4)
# Calculate the 95% confidence interval
confint(bootetasquared, level = 0.95, method = "quantile")
```
24. Based on this confidence interval, how much of an impact do the drug and obesity have on cholesterol?
*****
# Your turn
25. Let's analyze a dataset, `ToothGrowth` built-into R. It's results from a study on the effect of vitamin C on tooth growth in guinea pigs. The researchers were interested in the effects of two variables:
• `supp`: type of Vitamin C given to the guinea pigs (**OJ** = orange juice; **VC** = a liquid supplement)
• `dose`: the amount of Vitamin C given (**low**, **med**, or **high**).
This gives us a total of 6 treatment groups (*OJ-low, OJ-med, OJ-high, VC-low, VC-med, VC-high*). A total of 60 guinea pigs were randomly assigned, 10 to each of those groups.
Summary statistics and visualizations are displayed below.
**(a)** Evaluate the conditions necessary to conduct an AxB ANOVA. Are you concerned about any of the assumptions being violated?
**(b)** Based on the interaction plot, do you think we'll conclude that a significant interaction effect exists? Explain.
**(c)** Conduct an AxB (Supplement x Dose) ANOVA and draw a conclusion about the interaction effect.
**(d)** If the interaction effect is **not statistically significant**, then write a conclusion regarding the main effects of the type of supplement and dose. If the interaction effect **is statistically significant**, split your data by `dose` and test for differences between the **OJ** and **VC** treatments.
```{r}
# Load the dataset
data(ToothGrowth)
# Change the dose variable to a factor with levels: low, med, high
ToothGrowth$dose = factor(ToothGrowth$dose, levels=c(0.5,1.0,2.0),
labels=c("low","med","high"))
```
This table shows the means (and standard deviations) of the 10 guinea pigs in each combination of treatments.
. | Orange Juice | AbsorbAcid | Total
------:| ------------:| -------------:| ---------------:|
Low | 13.23 (4.460) | 07.98 (2.747) | **10.605 (4.4998)**
Med | 22.70 (3.911) | 16.77 (2.515) | **19.735 (4.4154)**
High | 26.06 (2.655) | 26.14 (4.798) | **26.100 (3.7742)**
**Total** | **20.66 (6.606)** | **16.96 (8.266)** | **N=60, 18.813 (7.6493)**
```{r}
# Density plot of each group
densityplot(~len | dose*supp, data=ToothGrowth,
main="Tooth Growth by Dose and Supplement",
xlab="Tooth Growth", layout=c(3,2))
# Side-by-side boxplots
bwplot(supp ~ len | dose, data=ToothGrowth,
ylab="supplement", xlab="Tooth Growth",
main="Tooth Growth by Dose and Supplement",
layout=(c(1,3)))
# Interaction plot
with(ToothGrowth, interaction.plot(x.factor=dose, trace.factor=supp,
response=len, fun=mean, type="b", legend=T,
ylab="mean tooth length", main="Interaction Plot",
pch=c(1,19)))
# Levene's test
leveneTest(len ~ supp * dose, data=ToothGrowth)
```
26. This time, analyze that same `ToothGrowth` dataset using a randomization-based equivalent to the AxB ANOVA. Test the interaction effect; then, split the data by dose and run randomization-based tests to compare pairs of means.
*****
# Publishing your solutions
When you're ready to try the **Your Turn** exercises:
- Download the appropriate [Your Turn file](http://bradthiessen.com/html5/M301.html).
- If the file downloads with a *.Rmd* extension, it should open automatically in RStudio.
- If the file downloads with a *.txt* extension (or if it does not open in RStudio), then:
- open the file in a text editor and copy its contents
- open RStudio and create a new **R Markdown** file (html file)
- paste the contents of the **Your Turn file** over the default contents of the R Markdown file.
- Type your solutions into the **Your Turn file**.
- The file attempts to show you *where* to type each of your solutions/answers.
- Once you're ready to publish your solutions, click the **Knit HTML** button located at the top of RStudio:  - Once you click that button, RStudio will begin working to create a .html file of your solutions.
- It may take a minute or two to compile everything, especially if your solutions contain simulation/randomization methods
- While the file is compiling, you may see some red text in the console.
- When the file is finished, it will open automatically.
- If the file does not open automatically, you'll see an error message in the console.
- Send me an [email](mailto:thiessenbradleya@sau.edu) with that error message and I should be able to help you out.
- Once you have a completed report of your solutions, look through it to see if everything looks ok. Then, send that yourturn.html file to me [via email](mailto:thiessenbradleya@sau.edu) or print it out and give it to me.
- Once you click that button, RStudio will begin working to create a .html file of your solutions.
- It may take a minute or two to compile everything, especially if your solutions contain simulation/randomization methods
- While the file is compiling, you may see some red text in the console.
- When the file is finished, it will open automatically.
- If the file does not open automatically, you'll see an error message in the console.
- Send me an [email](mailto:thiessenbradleya@sau.edu) with that error message and I should be able to help you out.
- Once you have a completed report of your solutions, look through it to see if everything looks ok. Then, send that yourturn.html file to me [via email](mailto:thiessenbradleya@sau.edu) or print it out and give it to me.
Sources:
This document is released under a [Creative Commons Attribution-ShareAlike 3.0 Unported](http://creativecommons.org/licenses/by-sa/3.0) license.

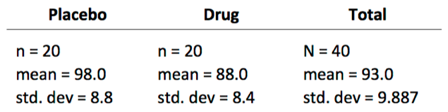
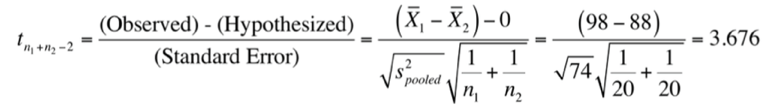
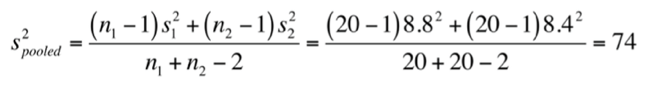






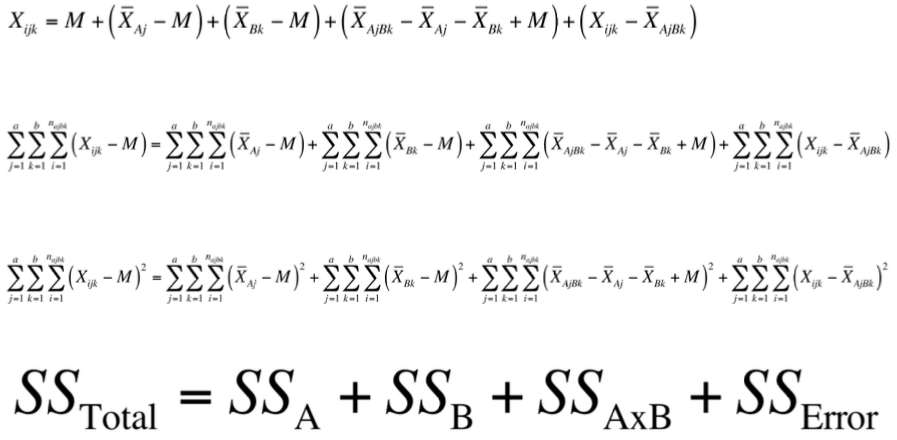





 - Once you click that button, RStudio will begin working to create a .html file of your solutions.
- It may take a minute or two to compile everything, especially if your solutions contain simulation/randomization methods
- While the file is compiling, you may see some red text in the console.
- When the file is finished, it will open automatically.
- If the file does not open automatically, you'll see an error message in the console.
- Send me an [email](mailto:thiessenbradleya@sau.edu) with that error message and I should be able to help you out.
- Once you have a completed report of your solutions, look through it to see if everything looks ok. Then, send that yourturn.html file to me [via email](mailto:thiessenbradleya@sau.edu) or print it out and give it to me.
- Once you click that button, RStudio will begin working to create a .html file of your solutions.
- It may take a minute or two to compile everything, especially if your solutions contain simulation/randomization methods
- While the file is compiling, you may see some red text in the console.
- When the file is finished, it will open automatically.
- If the file does not open automatically, you'll see an error message in the console.
- Send me an [email](mailto:thiessenbradleya@sau.edu) with that error message and I should be able to help you out.
- Once you have a completed report of your solutions, look through it to see if everything looks ok. Then, send that yourturn.html file to me [via email](mailto:thiessenbradleya@sau.edu) or print it out and give it to me.